What is High-Throughput Screening?
What is High-Throughput Screening?
High Throughput Screens (HTS) are recent scientific methods relevant to the field of chemistry and biology, in which hundreds of thousands of experimental samples are subjected to simultaneous testing under given conditions. The samples themselves may take the form of biochemical agents such as chemical compounds, amino acids, or live cells. With the development of laboratory robotics that automate sample preparation, handling and data analysis, scientists can easily and reliably generate and use large datasets from these HTS to answer complex biological questions. HTS is now widely used in the field of pharmaceuticals, biotech and academic institutes for drug discovery, target validation and the identification of genes or proteins that modulate a particular biological pathway.
Steps in a usual HTS experiment
There are multiple steps in any HTS experiment, which can take weeks to complete. However, these steps can be generalized into three categories: 1) Sample preparation, 2) sample handling, and 3) readouts and data acquisition.
Sample Preparation
HTS normally requires samples to be prepared in an arrayed format. The arrayed samples can be grown either on microtiter plates in liquid or on solid agar. The density of plates can range from 96, 192, 384, 768, 1,536, or 6,144. All these densities are multiples of 96, reflecting the original 96-well microtiter plate arranged in 8 x 12 with 9 mm spacing.
Library collections, most of which can be purchased commercially, are usually stored on 96 or 384 microtiter plates as stock plates. When needed, samples from stock plates are copied onto assay plates for HTS experiments.
Sample Handling: Liquid Handling
When doing Elisa-based anti-body screening or drug target validation, scientists often need to dispense precise volumes of liquid reagents. Dedicated liquid handling robots can precisely add and mix liquid reagents to multiple wells, which allows scientists to simultaneously screen for thousands of drugs, toxins, chemicals or bioactive compounds. It also allows scientists to dilute or sample liquid cultures in a highly precise manner (Figure 1).
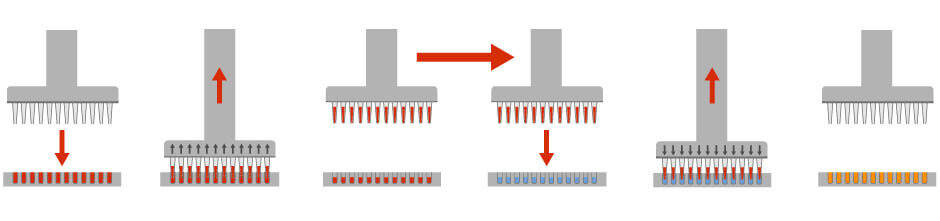
Figure 1: Liquid handling
Sample Handling: Colony Manipulation
Synthetic Genetic Array (SGA) and Yeast two-hybrid (Y2H) are examples of HTSs that involve the growth of microbial colonies on agar plates. In this scenario, it is necessary to pick, replicate, and re-array these colonies from one plate condition to the next (Figure 2). The ability to quickly and accurately manipulate colony arrays allows for the direct and unbiased comparison of any combination of mutant microorganisms.
The advantage of screening on agar is that the throughput can be much higher on agar than in liquid. Using the ROTOR+, scientists can screen more than 6,000 colonies on a single agar plate (1)!
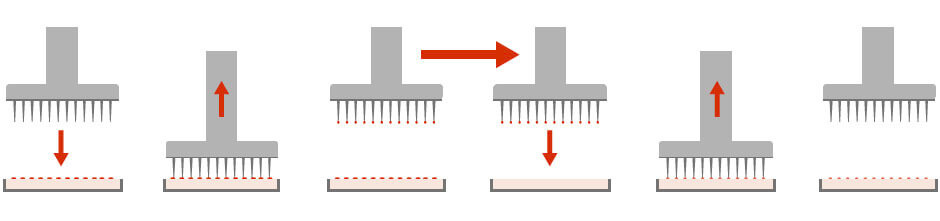
Figure 2: Colony picking
Additionally, ROTOR+ can be adapted to handle fickle organisms such as Chlamydomonas reinhardtii – which require high-level consistency for pinning to SGAs.
Chemical genetics is another HTS approach which can be used to identify potential therapeutic candidates. The method screens large numbers of colonies after treating them to large chemical libraries. Once treated the cell’s phenotype- behaviour and appearance is observed to identify chemicals with desirable effects like impaired growth to identify potential antibiotics and antifungals.
Chemical genetics have enabled Eric Brown at McMaster University to carry out phenotypic screens on Pseudomonas aeruginosa to identify synergistic combinations of compounds to treat the prevalent nosocomial pathogen.
Readouts & Data Acquisition
There are different readouts available for different HTS. Choosing the right readout depends on the balance between the cost of the screen, the desired data quality, and the biological questions that need to be answered.
Many HTS are interpreted through optical measurements – colour changes in cells and in liquid reactions, the turbidity of liquid culture, or fluorescence signals. For HTS that involve microbial colonies, colony size is often used (Figure 3). Data generated from these readouts are usually low in content richness because they are considered a proxy for a biological reaction or for an organism’s fitness. However, these forms of readouts are very cost-effective and are very useful for answering well-defined biological questions.
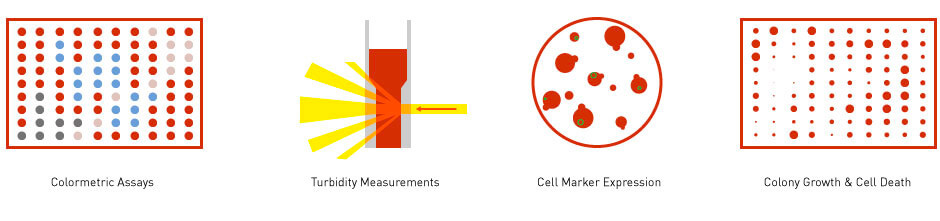
Figure 3: Readouts
High-content screens are any screening methods that document and analyze changes at the subcellular level across multiple conditions and time points. This form of readout is very costly to set up, requiring a sophisticated and expensive microscope or flow cytometry robotics. In return, high-content screens generate a large volume of content-rich information and allow multiple analyses to be applied.
Discover more applications of HTS
Dr Gianni Liti uses HTS experiments to understand genetic variation within the microbiome of oak trees.