Chemical Genetic Screening
The use of chemical genetics for drug target discovery
Summary
Chemical genetics is the study of genes through small-molecule perturbation. Chemical genetic screening is a phenotypic screening methodology that systematically tests the efficacy of thousands of small molecules simultaneously. It is superior to target-based screening methodology for drug discovery because chemical genetic screening is unbiased and screens chemicals directly in complex cellular environments. Therefore, chemical genetic screening is an efficient way to discover and validate new drugable targets and identify potentially efficacious therapeutics.
The Theory Behind Chemical Genetic Screening
Chemical probes are cell-permeable small molecules that potently inhibit protein function. Modern methods for identifying chemical probes rely on the screening of thousands of chemicals against a target(s) of interest in vitro (Figure 1A). However, this approach is ineffective, and the number of approved new drugs for treating human diseases has been in steady decline over the past decade. This is because in target-based screening where the target is pre-defined and drug-target interactions are conducted in vitro, the potency of the chemical in vitro rarely translates to low dosage and high specificity efficacy in vivo. It also limits our ability to discover other drug targets (1).
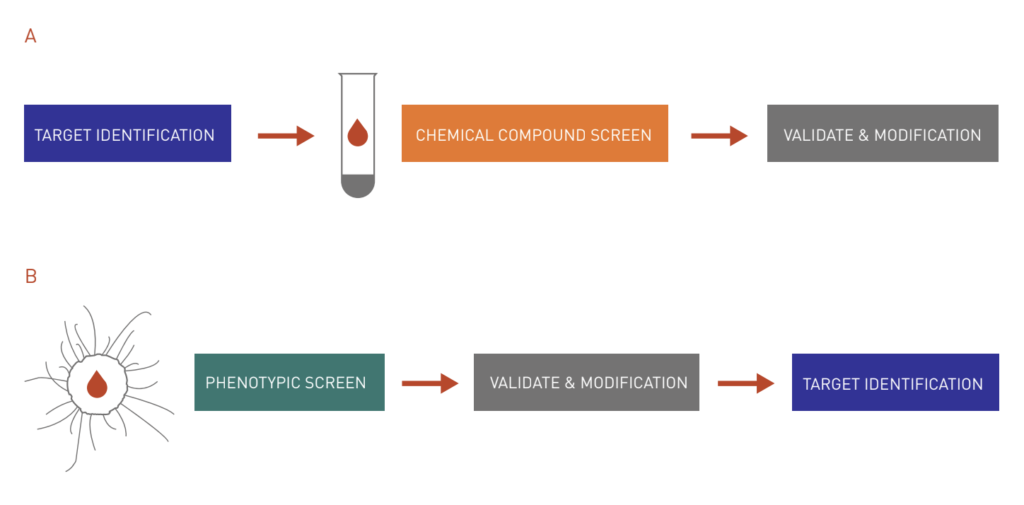
Figure 1: Target-based approach vs. chemical genetics approach to drug discovery.
A: The target-based approach begins with the selection of a protein target based on its relevance to a disease state. Then, a high-throughput screen is set up to look for chemicals that modulate target protein activities. Interesting hits are then validated and modified to improve their efficacy.
B: On the other hand, with the chemical genetics approach, a large chemical library is screened in a cell-based assay to find bioactive molecules for a disease state. After validation and modification, most interesting hits are subject to subsequent assays to identify cellular targets.
Phenotypic screening in model organisms is an alternative approach for discovering new small-molecule therapeutics that address the aforementioned shortfalls of current methodologies (Figure 1B). Phenotypic screening is a method for the systematic, unbiased, and parallel testing of thousands of molecules for a desired cellular phenotype. Employing cell-based phenotypic screening addresses two drawbacks specifically associated with conventional target-based screening methods. Firstly, screening using a cell-based assay directly measures drug potency in a complex cellular environment, which allows for the early assessment of the biological activities and off-target potential of a drug candidate. Secondly, screening for chemicals that alter the entire cellular pathway bypasses the bias of target pre-selection, leading to the unbiased discovery of new druggable targets (2).
Principles of chemical genetics in drug discovery
Two criteria must be met in a drug discovery process (3). Firstly, large chemical libraries must be screened to identify biologically active compounds that elicit the desired phenotype. Secondly, once the compounds are found, the relevant protein targets need to be identified. Chemical genetics can specifically address these two areas in drug discovery (Figure 2)
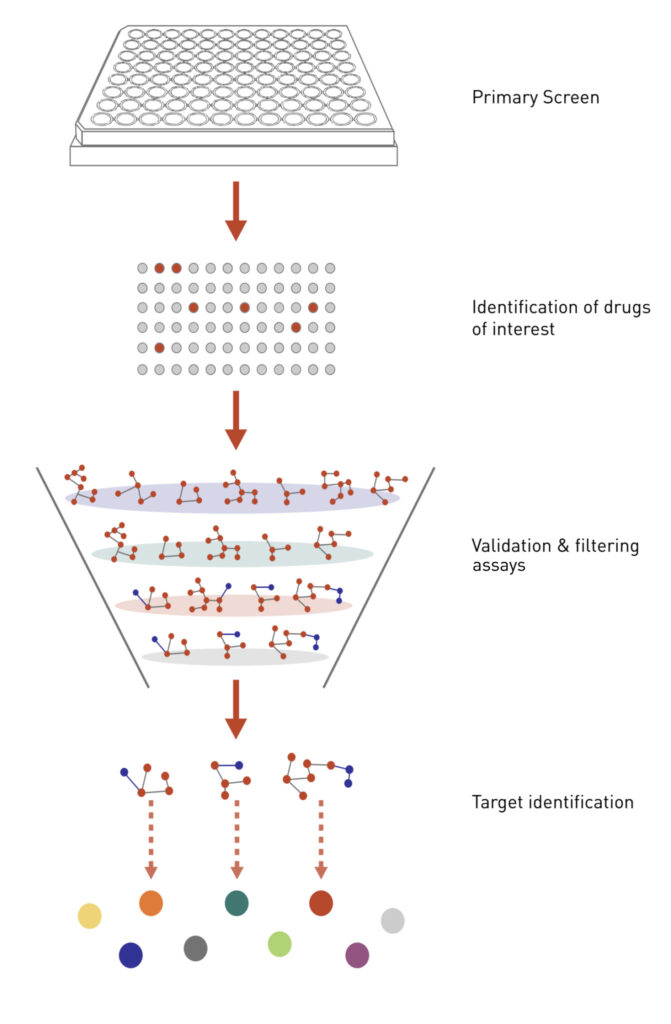
Figure 2: Applying chemical genetics to drug discovery.
A cell-based assay is established to screen for chemicals that induce a particular cellular phenotype. Once the hits from the primary screen are validated, the cellular targets of the most interesting hits are then identified using various genetic assays.
Compound screening
In a typical screen to identify bioactive compounds, a diverse library of compounds is applied to the cells that mimic a particular disease state, causing perturbations to different cellular pathways and resulting in phenotypic responses. Compounds that specifically elicit desired phenotypes will be identified and subjected to follow-up studies to identify their protein target (4).
Target Identification
Genetic screening is an effective strategy for target identification, especially in organisms where whole genome library collections are available. There are three gene-dosage based assays in yeast that are commonly used to identify drug targets. All three assays are unbiased growth-based assays that do not require prior knowledge of a compound’s mechanism of action (3,5).
HaploInsufficiency Profiling (HIP) assay is based on the principle that a decreased dosage of a drug target gene can result in an increase in drug sensitivity (Figure 3A). In a HIP assay, heterozygous deletion mutants are treated with a drug of interest. Strains that exhibit increased sensitivity in the form of growth inhibition will be identified. The HIP assay reveals the direct target as well as other components in the same pathway.
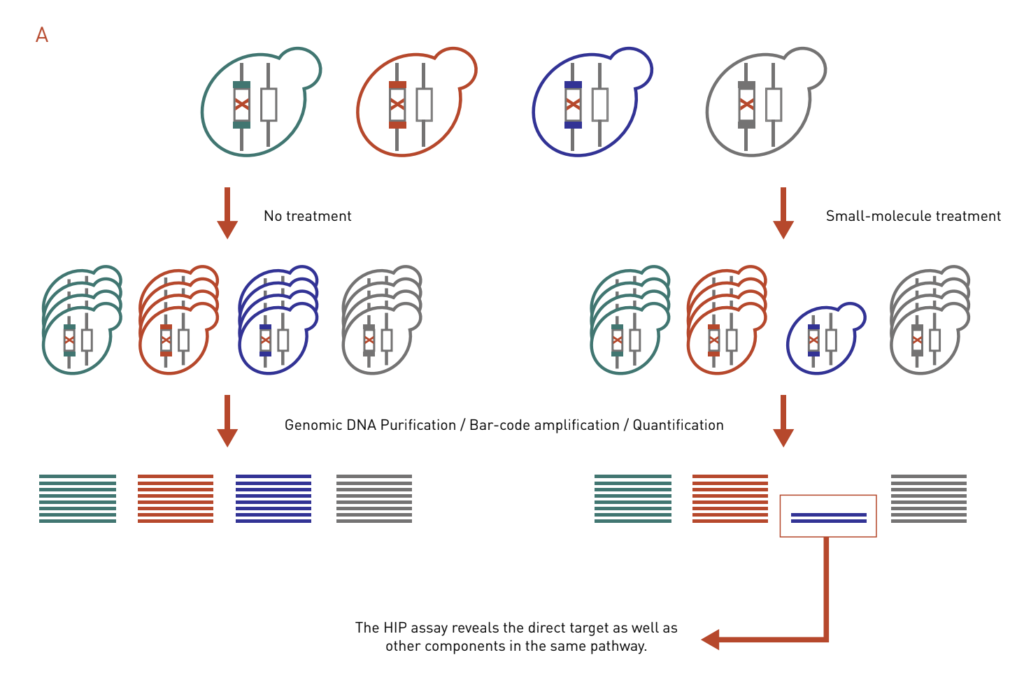
Figure 3: Chemical genetics-based dosage-dependent assays for target identification. A pool of molecular barcoded yeast strains is grown in the presence or absence of chemical compounds. At the end of the experiment, the barcodes are purified, amplified, and quantified to give the relative abundance of each strain in the pool.
A: The HIP assay uses heterozygous deletion collection to look for strains that are sensitive to drug treatment, which reveals direct targets and other genes in the same pathway.
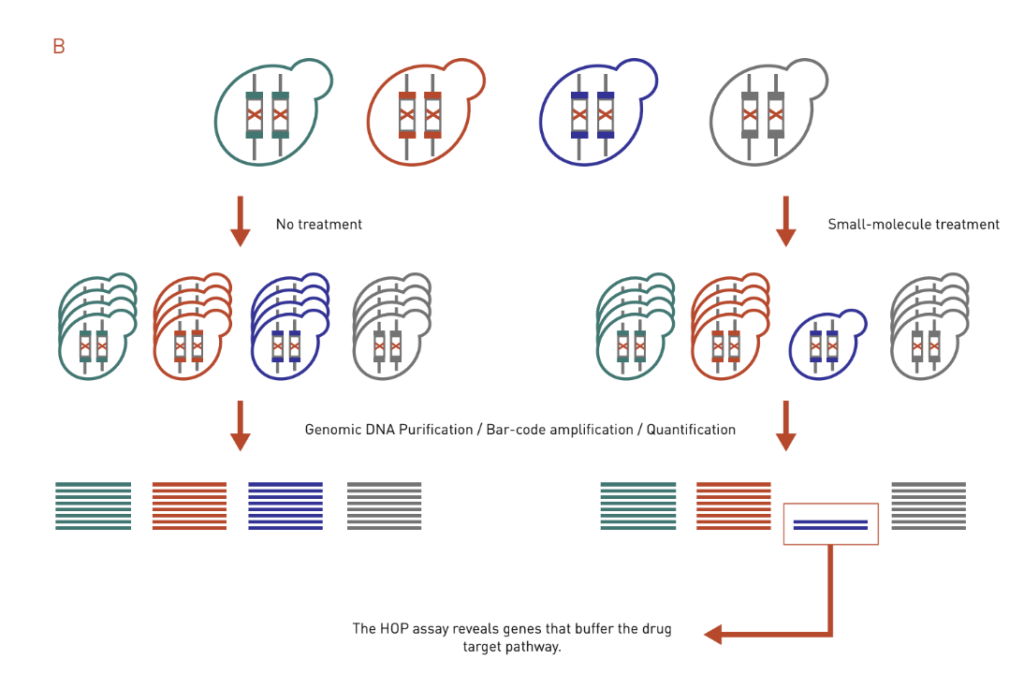
B: The HOP assay works similarly to the HIP assay. It uses ho-mozygous deletion collection and identifies genes that buffer the drug target pathway.
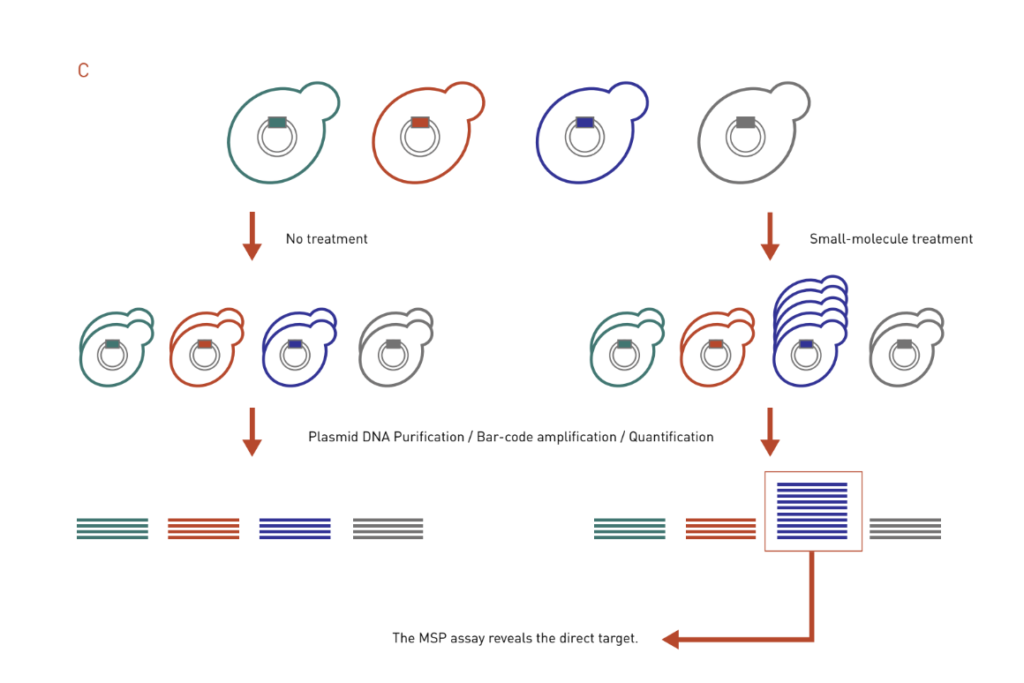
C: The MSP works in the opposite way of the HIP assay. It uses the over-expression DNA library to look for strains that are resistant to drug treatment, thereby identifying direct targets of the drug.
Homozygous Profiling (HOP) assay is similar to the HIP assay except that both copies of a nonessential gene are deleted (Figure 3B). Instead of identifying a direct target, the HOP assay often identifies genes that buffer the drug target pathway. This is because the HOP assay mimics a double deletion mutant in that the second gene disruption is achieved via compound inhibition. Direct target identification, however, can be inferred by integrating HOP chemical-genetic profiles with genetic interaction profile. The chemical-genetic profile of a drug will be similar to the genetic interaction profile of its target gene.
Multicopy suppression profiling (MSP) assay is based on the principle that increased dosage of a drug target gene can confer resistance to drug-mediated growth inhibition, essentially a mirror to the HIP assay (Figure 3C). In a MSP assay, strains that contain high-copy overexpression plasmids are treated with a drug of interest, and individuals that exhibit growth advantage in the presence of the drug will be identified. Like the HIP assay, MSP assay often identifies direct target of a drug.
All three gene-dosage-based screening assays are often performed in a single liquid culture using bar-coded strains to assess the competitive growth of the entire collection6. However, these assays can also be performed on solid media using colony size to measure individual colony fitness7
S. Cerevisiae as a Platform for Chemical Genetic Screening and Drug Identification
S. cerevisiae is ideally suited for high-throughput phenotypic screening for several reasons. Firstly, it has a short doubling time and simple growth requirements. Secondly, its genome is the best characterized among eukaryotic models. Thirdly, many core cellular processes in S. cerevisiae, such as cell cycle control, DNA repair and various metabolic pathways, are conserved in humans (8,9). Finally, many laboratories have experience using yeast strains to model disease state in human cells and screen for promising compounds (4).
However, it usually requires a much higher dosage of chemicals to perform genetic screens in yeast than in mammalian cells. This is because yeast has a cell wall, making it hard for some chemicals to enter the cells 10. In addition, yeast has a dynamic chemical defence, which uses efflux pumps that actively transport drugs out of the cells (11,12). Special yeast strains that are mutated with these genes will increase the drug sensitivity of a yeast cell (13).
Library Small Compound Libraries
In a forward chemical genetic screen, it is desirable to screen for a selection of chemicals that cover as much chemical space as possible. However, since the chemical space of all possible chemical structures is extremely large, and the screen itself is compound-intensive and time-consuming, it is very important to pre-select a subset of compounds that are enriched for known active substructures or most likely to accumulate in the organism of interest (14). Many public and private institutes hold large collections of small molecule libraries for drug discovery and chemical genetics screenings (15-19).
Working with Libraries
High-throughput chemical genomics screening can be done quickly and easily using automated robotics, such as the Singer ROTOR+, which can rapidly pin high-density arrays of bacteria and yeast colonies. In addition, the ROTOR+ screening robot provides an automated platform to simultaneously produce multiple experimental replicates.

By Jay Yang, PhD | Former Scientific Advisor at Singer Instruments
Looking for screening tools to accelerate your research?
ROTOR+: the only microbial screening solution capable
of pinning a million colonies an hour.